Visualization
 "Visualization is any technique for creating images, diagrams, or animations to communicate a message. Visualization through visual imagery has been an effective way to communicate both abstract and concrete ideas since the dawn of man. Examples from history include cave paintings, Egyptian hieroglyphs, Greek geometry, and Leonardo da Vinci's revolutionary methods of technical drawing for engineering and scientific purposes." (Wikipedia) Our own research focus is the 3D Visualization, meaning the visualization of three dimensional objects. Example applications using 3D Visualization are:
"Visualization is any technique for creating images, diagrams, or animations to communicate a message. Visualization through visual imagery has been an effective way to communicate both abstract and concrete ideas since the dawn of man. Examples from history include cave paintings, Egyptian hieroglyphs, Greek geometry, and Leonardo da Vinci's revolutionary methods of technical drawing for engineering and scientific purposes." (Wikipedia) Our own research focus is the 3D Visualization, meaning the visualization of three dimensional objects. Example applications using 3D Visualization are:
- Computer Aided Construction (architecture, engineering, ...)
- the visualization of 3-dimensional medical data, for example from computer tomography,
- the visualization of landscape data, showing either status quo or future measurements.
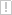
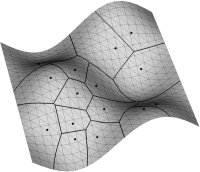
Visualization is fundamental for many geometrical problems. Whenever a new algorithm for the calculation of a 3D Voronoi Diagram or medial axis is implemented, the very first test if the new method produces the right results is usually optical.
One of the first projects of the Welfenlab was the developement of a NURBS base visualization software called "Janus" ("Just Another NUrbs System"). Despite his name, the program could visualize not only NURBS surfaces but also bezier patches (and curves), triangle surfaces, point clouds or curve-on-surface-models.
3D Visualization in the Field of Medicine
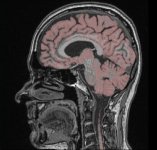
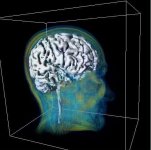
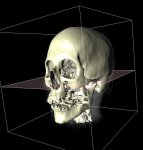
A currently very hot focus of our department is the visualization of voxel data in the field of medicine. A voxel can be seen as a three dimensional analogon of a pixel. A pixel (defined by a x-, y- and a color value) describes a point on a computer monitor, while the voxel (which contains an additional z-value) describes a three dimensional point in space. Voxel data in a medical context is gained by different imaging techniques, such as computer tomography (CT) or magnetic resonance imaging (MRI). The question remains, how such voxel data can be visualized in a human-understandable, natural way. Depending on the application it is either necessary to produce high quality images or a fast, interactive form of visualization where the user can move free around (or through) the virtual object.
 |  |  Â Â |
Visualization of medical voxel data is often combined with the problem of segmentation, which describes the partitioning of our (three dimensional) image into multiple regions. As an exampel, a part of a voxel data set gained by a CT scan might be described as "bone", a subset of the bone might be called "jaw". Both, "bone" and "jaw" are called segments, the process of their creation (manual or automatic) is called segmentation.
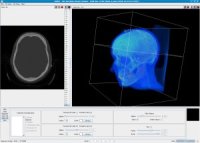 Both applications, visualization and segmentation of medical data is in the focus of our research. For this purpose, we developed the sofware YaDiV ("Yet Another Dicom Viewer"), which is able to visualize, segment and register medical data (DICOM directory format). The software is currently used as a platform for our own research and is evaluated with medical scientists from the Medizinischen Hochschule Hannover (MHH). There exist concrete plans to publish YaDiV as open source in the next weeks. More about YaDiV ...
Both applications, visualization and segmentation of medical data is in the focus of our research. For this purpose, we developed the sofware YaDiV ("Yet Another Dicom Viewer"), which is able to visualize, segment and register medical data (DICOM directory format). The software is currently used as a platform for our own research and is evaluated with medical scientists from the Medizinischen Hochschule Hannover (MHH). There exist concrete plans to publish YaDiV as open source in the next weeks. More about YaDiV ...
Visualization with Game Engines
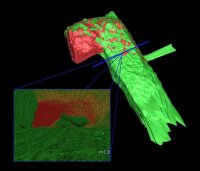 Another focus of our visualization research is the so called "serious gaming". In the recent years, the computer gaming industry has become a large and important market and impressive amounts of money are spent on the development of new game engines. In contrast to their development costs, the price for the final product is very low compared to a professional 3D visualization/animation program.
Another focus of our visualization research is the so called "serious gaming". In the recent years, the computer gaming industry has become a large and important market and impressive amounts of money are spent on the development of new game engines. In contrast to their development costs, the price for the final product is very low compared to a professional 3D visualization/animation program.
The idea to use this potential for other purposes than gaming seems obvious.
Arbeiten
- bachelor thesis Adaptive 3D-Visualization of fractal Heightmaps using Tessellation Shaders. Adrian Hengelhaupt - 03/2014
- masters thesis Implementation und Analyse von Triangle Strip Algorithmen. Friedrich Hattendorf - 05/2013
- bachelor thesis Vegetationsgenerierung mittels Zellulärer Automaten und Spieltheorie. Sven Röttering - 02/2013
- bachelor thesis Stereographische Visualisierung von 360° Aufnahmen auf einem Head Mounted Display (HMD). Peter Denis - 09/2012
- bachelor thesis Implementation eines Echtzeitmodells fĂĽr den Wasserkreislauf. Oliver Weidner - 09/2012
- bachelor thesis Prozedurale Multiskalen-Generierung von Planetenoberflächen. Felix Haase - 06/2012
- diploma thesis Halbautomatische Segmentierung der knöchernen Strukturen des Kniegelenks in MRT Aufnahmen. Benjamin Fleischer - 05/2012
- bachelor thesis 3D-Textursynthese auf Grundlage zweidimensionaler Exemplare. Romeo Shuka - 05/2012
- Studienarbeit Dynamische Erzeungung von Triangle Strips als Erweiterung des Marching Cube Verfahrens. Friedrich KrĂĽger - 12/2011
- bachelor thesis Modelling of realistic Blood Vessel Geometry. Sergi Lazaro - 09/2011
- Studienarbeit Extraktion von Isoflächen mit Hilfe des Marching-Cubes-Algorithmus. Damaris Schumann - 04/2009
- masters thesis High-Quality Ray Casting Methods for the Visualization of segmented medical voxel data. Maximilian MĂĽller - 04/2009
- Studienarbeit Untersuchung zur Grauwertdarstellung auf LC-Displays. Lara Toma - 06/2008
- bachelor thesis OpenSG - Verteilte Visualisierung in verschiedenen Anwendungsszenarien. Marcus Rieche - 06/2007
- masters thesis Dynamische 3D-Visualisierung eines Radial-Axial-Ringwalzprozesses. Maxim Marchenko - 12/2006
- masters thesis Untersuchungen zur Eignung von 3D-Spieleengines fĂĽr die wissenschaftliche Visualisierung anhand der CryENGINE. Marc Herrlich - 03/2006
- Studienarbeit Non photorealistic rendering: Schnittflächenverfahren und Implementierung. Yifan Yu - 02/2006
- bachelor thesis 3D-Visualisierung mit Hilfe von Spiel-Engines unter Berücksichtigung erweiterter Texturierungsmöglichkeiten. Michael Hanel - 12/2004
- Studienarbeit Portierung des Visualisierungssystems Janus nach Linux unter besonderer BerĂĽcksichtigung objektorientierter Signale. Jennis Meyer-Spradow - 12/2003
- Studienarbeit Illustrative Volume Rendering with Lit Sphere Mapping using additional Segment Information. Johannes Wahle - in Bearbeitung
- Studienarbeit CellARS: Cell Architecture Radar Simulation. Daniel Brandes - 10/2010


